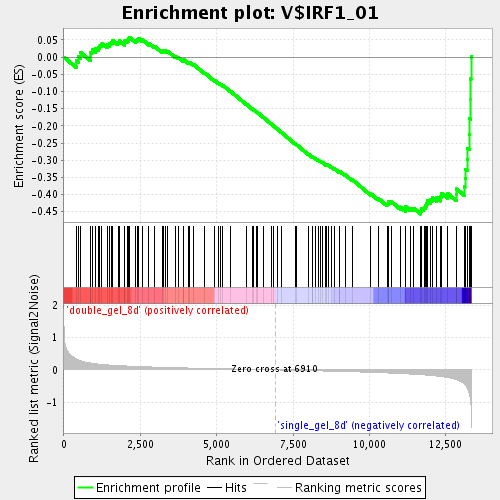
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_single_gel_8d.class.cls #double_gel_8d_versus_single_gel_8d_repos |
| Phenotype | class.cls#double_gel_8d_versus_single_gel_8d_repos |
| Upregulated in class | single_gel_8d |
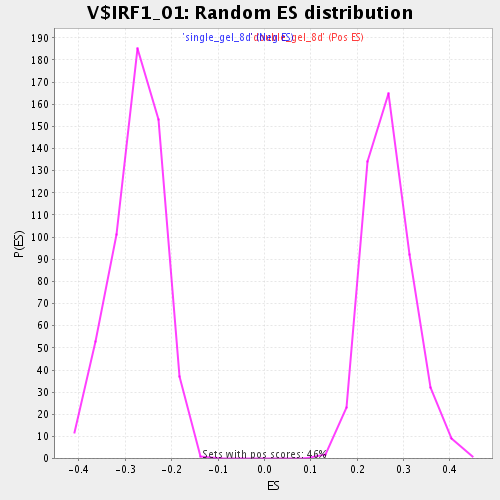
| GeneSet | V$IRF1_01 |
| Enrichment Score (ES) | -0.45678362 |
| Normalized Enrichment Score (NES) | -1.6617578 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.09212872 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | EGR2 | EGR2 Entrez, Source | early growth response 2 (Krox-20 homolog, Drosophila) | 402 | 0.338 | -0.0105 | No |
| 2 | PEX12 | PEX12 Entrez, Source | peroxisomal biogenesis factor 12 | 486 | 0.308 | 0.0013 | No |
| 3 | KYNU | KYNU Entrez, Source | kynureninase (L-kynurenine hydrolase) | 548 | 0.283 | 0.0133 | No |
| 4 | EPAS1 | EPAS1 Entrez, Source | endothelial PAS domain protein 1 | 871 | 0.211 | 0.0014 | No |
| 5 | BCL11A | BCL11A Entrez, Source | B-cell CLL/lymphoma 11A (zinc finger protein) | 875 | 0.211 | 0.0135 | No |
| 6 | EHD1 | EHD1 Entrez, Source | EH-domain containing 1 | 920 | 0.204 | 0.0221 | No |
| 7 | ASPA | ASPA Entrez, Source | aspartoacylase (Canavan disease) | 1027 | 0.189 | 0.0253 | No |
| 8 | STX17 | STX17 Entrez, Source | syntaxin 17 | 1124 | 0.180 | 0.0286 | No |
| 9 | BAT5 | BAT5 Entrez, Source | HLA-B associated transcript 5 | 1179 | 0.173 | 0.0346 | No |
| 10 | MGAT2 | MGAT2 Entrez, Source | mannosyl (alpha-1,6-)-glycoprotein beta-1,2-N-acetylglucosaminyltransferase | 1244 | 0.168 | 0.0396 | No |
| 11 | ATF3 | ATF3 Entrez, Source | activating transcription factor 3 | 1414 | 0.156 | 0.0360 | No |
| 12 | EGFL6 | EGFL6 Entrez, Source | EGF-like-domain, multiple 6 | 1486 | 0.151 | 0.0395 | No |
| 13 | DLX1 | DLX1 Entrez, Source | distal-less homeobox 1 | 1542 | 0.146 | 0.0439 | No |
| 14 | PITX2 | PITX2 Entrez, Source | paired-like homeodomain transcription factor 2 | 1599 | 0.143 | 0.0481 | No |
| 15 | PPARGC1A | PPARGC1A Entrez, Source | peroxisome proliferative activated receptor, gamma, coactivator 1, alpha | 1770 | 0.133 | 0.0431 | No |
| 16 | BST2 | BST2 Entrez, Source | bone marrow stromal cell antigen 2 | 1811 | 0.130 | 0.0477 | No |
| 17 | LMO4 | LMO4 Entrez, Source | LIM domain only 4 | 1984 | 0.122 | 0.0419 | No |
| 18 | LMO3 | LMO3 Entrez, Source | LIM domain only 3 (rhombotin-like 2) | 1995 | 0.122 | 0.0483 | No |
| 19 | CDK6 | CDK6 Entrez, Source | cyclin-dependent kinase 6 | 2067 | 0.119 | 0.0499 | No |
| 20 | BMP4 | BMP4 Entrez, Source | bone morphogenetic protein 4 | 2116 | 0.117 | 0.0531 | No |
| 21 | DMD | DMD Entrez, Source | dystrophin (muscular dystrophy, Duchenne and Becker types) | 2146 | 0.116 | 0.0578 | No |
| 22 | LIF | LIF Entrez, Source | leukemia inhibitory factor (cholinergic differentiation factor) | 2346 | 0.109 | 0.0491 | No |
| 23 | ZPBP2 | ZPBP2 Entrez, Source | zona pellucida binding protein 2 | 2397 | 0.107 | 0.0516 | No |
| 24 | ARHGAP5 | ARHGAP5 Entrez, Source | Rho GTPase activating protein 5 | 2435 | 0.105 | 0.0550 | No |
| 25 | CGA | CGA Entrez, Source | glycoprotein hormones, alpha polypeptide | 2568 | 0.100 | 0.0509 | No |
| 26 | CCL5 | CCL5 Entrez, Source | chemokine (C-C motif) ligand 5 | 2781 | 0.092 | 0.0403 | No |
| 27 | PLEKHB1 | PLEKHB1 Entrez, Source | pleckstrin homology domain containing, family B (evectins) member 1 | 2957 | 0.086 | 0.0322 | No |
| 28 | UBD | UBD Entrez, Source | ubiquitin D | 3213 | 0.078 | 0.0175 | No |
| 29 | GLRA1 | GLRA1 Entrez, Source | glycine receptor, alpha 1 (startle disease/hyperekplexia, stiff man syndrome) | 3262 | 0.077 | 0.0184 | No |
| 30 | IL6 | IL6 Entrez, Source | interleukin 6 (interferon, beta 2) | 3320 | 0.076 | 0.0185 | No |
| 31 | FBXO8 | FBXO8 Entrez, Source | F-box protein 8 | 3404 | 0.074 | 0.0165 | No |
| 32 | CACNB2 | CACNB2 Entrez, Source | calcium channel, voltage-dependent, beta 2 subunit | 3644 | 0.068 | 0.0025 | No |
| 33 | NOG | NOG Entrez, Source | noggin | 3747 | 0.065 | -0.0014 | No |
| 34 | ATP1B4 | ATP1B4 Entrez, Source | ATPase, (Na+)/K+ transporting, beta 4 polypeptide | 3912 | 0.062 | -0.0102 | No |
| 35 | CREB1 | CREB1 Entrez, Source | cAMP responsive element binding protein 1 | 3915 | 0.062 | -0.0067 | No |
| 36 | PRKCB1 | PRKCB1 Entrez, Source | protein kinase C, beta 1 | 4062 | 0.058 | -0.0143 | No |
| 37 | DGKB | DGKB Entrez, Source | diacylglycerol kinase, beta 90kDa | 4120 | 0.057 | -0.0153 | No |
| 38 | DNASE1L3 | DNASE1L3 Entrez, Source | deoxyribonuclease I-like 3 | 4230 | 0.054 | -0.0204 | No |
| 39 | IFNB1 | IFNB1 Entrez, Source | interferon, beta 1, fibroblast | 4609 | 0.046 | -0.0462 | No |
| 40 | MXD3 | MXD3 Entrez, Source | MAX dimerization protein 3 | 4925 | 0.039 | -0.0677 | No |
| 41 | PRDX5 | PRDX5 Entrez, Source | peroxiredoxin 5 | 5059 | 0.037 | -0.0756 | No |
| 42 | CXCL10 | CXCL10 Entrez, Source | chemokine (C-X-C motif) ligand 10 | 5128 | 0.035 | -0.0786 | No |
| 43 | PITX1 | PITX1 Entrez, Source | paired-like homeodomain transcription factor 1 | 5205 | 0.034 | -0.0824 | No |
| 44 | BLK | BLK Entrez, Source | B lymphoid tyrosine kinase | 5455 | 0.029 | -0.0995 | No |
| 45 | LGI1 | LGI1 Entrez, Source | leucine-rich, glioma inactivated 1 | 5982 | 0.019 | -0.1381 | No |
| 46 | VAMP3 | VAMP3 Entrez, Source | vesicle-associated membrane protein 3 (cellubrevin) | 6158 | 0.015 | -0.1505 | No |
| 47 | PTPRR | PTPRR Entrez, Source | protein tyrosine phosphatase, receptor type, R | 6189 | 0.014 | -0.1519 | No |
| 48 | PAX3 | PAX3 Entrez, Source | paired box gene 3 (Waardenburg syndrome 1) | 6318 | 0.012 | -0.1609 | No |
| 49 | ACCN1 | ACCN1 Entrez, Source | amiloride-sensitive cation channel 1, neuronal (degenerin) | 6344 | 0.011 | -0.1621 | No |
| 50 | RIMS2 | RIMS2 Entrez, Source | regulating synaptic membrane exocytosis 2 | 6544 | 0.007 | -0.1768 | No |
| 51 | IL11 | IL11 Entrez, Source | interleukin 11 | 6783 | 0.003 | -0.1946 | No |
| 52 | GNAO1 | GNAO1 Entrez, Source | guanine nucleotide binding protein (G protein), alpha activating activity polypeptide O | 6866 | 0.001 | -0.2007 | No |
| 53 | DIO2 | DIO2 Entrez, Source | deiodinase, iodothyronine, type II | 6980 | -0.002 | -0.2092 | No |
| 54 | TAPBP | TAPBP Entrez, Source | TAP binding protein (tapasin) | 7131 | -0.005 | -0.2202 | No |
| 55 | SHANK2 | SHANK2 Entrez, Source | SH3 and multiple ankyrin repeat domains 2 | 7570 | -0.014 | -0.2525 | No |
| 56 | LTBP2 | LTBP2 Entrez, Source | latent transforming growth factor beta binding protein 2 | 7613 | -0.015 | -0.2548 | No |
| 57 | DLG1 | DLG1 Entrez, Source | discs, large homolog 1 (Drosophila) | 8008 | -0.024 | -0.2832 | No |
| 58 | CSAD | CSAD Entrez, Source | cysteine sulfinic acid decarboxylase | 8121 | -0.026 | -0.2901 | No |
| 59 | TFEC | TFEC Entrez, Source | transcription factor EC | 8147 | -0.026 | -0.2905 | No |
| 60 | HPCAL1 | HPCAL1 Entrez, Source | hippocalcin-like 1 | 8231 | -0.028 | -0.2951 | No |
| 61 | SFRS12 | SFRS12 Entrez, Source | splicing factor, arginine/serine-rich 12 | 8323 | -0.030 | -0.3002 | No |
| 62 | AXIN1 | AXIN1 Entrez, Source | axin 1 | 8385 | -0.032 | -0.3029 | No |
| 63 | ESR1 | ESR1 Entrez, Source | estrogen receptor 1 | 8446 | -0.033 | -0.3055 | No |
| 64 | PSMB10 | PSMB10 Entrez, Source | proteasome (prosome, macropain) subunit, beta type, 10 | 8574 | -0.036 | -0.3129 | No |
| 65 | TWIST1 | TWIST1 Entrez, Source | twist homolog 1 (acrocephalosyndactyly 3; Saethre-Chotzen syndrome) (Drosophila) | 8609 | -0.037 | -0.3133 | No |
| 66 | B2M | B2M Entrez, Source | beta-2-microglobulin | 8650 | -0.038 | -0.3141 | No |
| 67 | ACIN1 | ACIN1 Entrez, Source | apoptotic chromatin condensation inducer 1 | 8761 | -0.040 | -0.3201 | No |
| 68 | MAP3K11 | MAP3K11 Entrez, Source | mitogen-activated protein kinase kinase kinase 11 | 8842 | -0.042 | -0.3237 | No |
| 69 | ROCK1 | ROCK1 Entrez, Source | Rho-associated, coiled-coil containing protein kinase 1 | 9002 | -0.046 | -0.3330 | No |
| 70 | VAX1 | VAX1 Entrez, Source | ventral anterior homeobox 1 | 9033 | -0.047 | -0.3325 | No |
| 71 | KARS | KARS Entrez, Source | lysyl-tRNA synthetase | 9206 | -0.051 | -0.3425 | No |
| 72 | GATA3 | GATA3 Entrez, Source | GATA binding protein 3 | 9441 | -0.058 | -0.3568 | No |
| 73 | NPR3 | NPR3 Entrez, Source | natriuretic peptide receptor C/guanylate cyclase C (atrionatriuretic peptide receptor C) | 10036 | -0.076 | -0.3972 | No |
| 74 | NR3C2 | NR3C2 Entrez, Source | nuclear receptor subfamily 3, group C, member 2 | 10284 | -0.084 | -0.4109 | No |
| 75 | KPNA3 | KPNA3 Entrez, Source | karyopherin alpha 3 (importin alpha 4) | 10575 | -0.094 | -0.4273 | No |
| 76 | ITPKB | ITPKB Entrez, Source | inositol 1,4,5-trisphosphate 3-kinase B | 10615 | -0.096 | -0.4246 | No |
| 77 | YWHAG | YWHAG Entrez, Source | tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, gamma polypeptide | 10619 | -0.096 | -0.4192 | No |
| 78 | MXI1 | MXI1 Entrez, Source | MAX interactor 1 | 10722 | -0.099 | -0.4211 | No |
| 79 | NFIX | NFIX Entrez, Source | nuclear factor I/X (CCAAT-binding transcription factor) | 11009 | -0.112 | -0.4361 | No |
| 80 | DOCK9 | DOCK9 Entrez, Source | dedicator of cytokinesis 9 | 11177 | -0.120 | -0.4417 | No |
| 81 | CNTF | CNTF Entrez, Source | ciliary neurotrophic factor | 11187 | -0.121 | -0.4353 | No |
| 82 | USF1 | USF1 Entrez, Source | upstream transcription factor 1 | 11338 | -0.128 | -0.4391 | No |
| 83 | SLC22A17 | SLC22A17 Entrez, Source | solute carrier family 22 (organic cation transporter), member 17 | 11450 | -0.135 | -0.4396 | No |
| 84 | EIF4A2 | EIF4A2 Entrez, Source | eukaryotic translation initiation factor 4A, isoform 2 | 11679 | -0.148 | -0.4481 | Yes |
| 85 | BLNK | BLNK Entrez, Source | B-cell linker | 11704 | -0.150 | -0.4411 | Yes |
| 86 | PSMB8 | PSMB8 Entrez, Source | proteasome (prosome, macropain) subunit, beta type, 8 (large multifunctional peptidase 7) | 11784 | -0.155 | -0.4379 | Yes |
| 87 | PSMB9 | PSMB9 Entrez, Source | proteasome (prosome, macropain) subunit, beta type, 9 (large multifunctional peptidase 2) | 11832 | -0.159 | -0.4322 | Yes |
| 88 | TERF2IP | TERF2IP Entrez, Source | telomeric repeat binding factor 2, interacting protein | 11858 | -0.161 | -0.4246 | Yes |
| 89 | FOXA1 | FOXA1 Entrez, Source | forkhead box A1 | 11900 | -0.165 | -0.4180 | Yes |
| 90 | NRAS | NRAS Entrez, Source | neuroblastoma RAS viral (v-ras) oncogene homolog | 11999 | -0.173 | -0.4153 | Yes |
| 91 | SMAD6 | SMAD6 Entrez, Source | SMAD, mothers against DPP homolog 6 (Drosophila) | 12050 | -0.177 | -0.4086 | Yes |
| 92 | TAP1 | TAP1 Entrez, Source | transporter 1, ATP-binding cassette, sub-family B (MDR/TAP) | 12194 | -0.191 | -0.4082 | Yes |
| 93 | DDIT3 | DDIT3 Entrez, Source | DNA-damage-inducible transcript 3 | 12335 | -0.209 | -0.4065 | Yes |
| 94 | VGLL4 | VGLL4 Entrez, Source | vestigial like 4 (Drosophila) | 12356 | -0.210 | -0.3957 | Yes |
| 95 | USP18 | USP18 Entrez, Source | ubiquitin specific peptidase 18 | 12554 | -0.237 | -0.3967 | Yes |
| 96 | CXCL11 | CXCL11 Entrez, Source | chemokine (C-X-C motif) ligand 11 | 12838 | -0.299 | -0.4005 | Yes |
| 97 | GATA6 | GATA6 Entrez, Source | GATA binding protein 6 | 12847 | -0.302 | -0.3834 | Yes |
| 98 | CDKN2A | CDKN2A Entrez, Source | cyclin-dependent kinase inhibitor 2A (melanoma, p16, inhibits CDK4) | 13099 | -0.425 | -0.3774 | Yes |
| 99 | NGFRAP1 | NGFRAP1 Entrez, Source | nerve growth factor receptor (TNFRSF16) associated protein 1 | 13127 | -0.447 | -0.3532 | Yes |
| 100 | FXYD5 | FXYD5 Entrez, Source | FXYD domain containing ion transport regulator 5 | 13147 | -0.472 | -0.3269 | Yes |
| 101 | STX11 | STX11 Entrez, Source | syntaxin 11 | 13195 | -0.556 | -0.2978 | Yes |
| 102 | NDN | NDN Entrez, Source | necdin homolog (mouse) | 13215 | -0.586 | -0.2649 | Yes |
| 103 | COL4A1 | COL4A1 Entrez, Source | collagen, type IV, alpha 1 | 13269 | -0.760 | -0.2242 | Yes |
| 104 | TGFB3 | TGFB3 Entrez, Source | transforming growth factor, beta 3 | 13277 | -0.783 | -0.1788 | Yes |
| 105 | LOX | LOX Entrez, Source | lysyl oxidase | 13308 | -0.991 | -0.1228 | Yes |
| 106 | PDGFC | PDGFC Entrez, Source | platelet derived growth factor C | 13311 | -1.014 | -0.0634 | Yes |
| 107 | F3 | F3 Entrez, Source | coagulation factor III (thromboplastin, tissue factor) | 13324 | -1.119 | 0.0014 | Yes |