Download source code implementing our Linker algorithm. Ensure that the following 3rd party dependencies are installed on your machine:
The supplementary input files directory contains the weighted, directed yeast interactome, the target nodes, and the list of queries used as inputs to Linker for analyzing the Chen2004 yeast cell cycle model:
1.) Suppose we want to link the protein YMR001C (Cdc5) to the proteins in the
Chen2004 model of the yeast cell cycle. First, create an input query file:
echo YMR001C > query-YMR001C.txt
Now, we can ask Linker to identify the most probable paths connecting YMR001C to
Chen2004 as follows:
python linker.py yeast-interactome.txt query-YMR001C.txt chen2004-species.txt -o cdc5
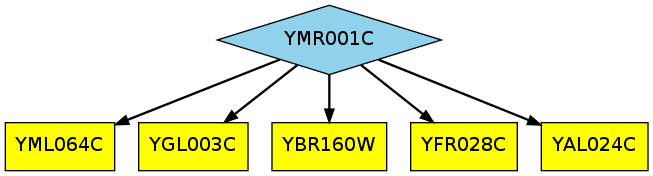
By default, Linker will compute the k=5 most probable paths linking
YMR001C to the cell cycle, and you should have the following three output
files:

2.) The outputs may be difficult to interpret, since the identifiers are in the
"systematic" namespace (e.g., YMR001C). We can use the -m
option to map the nodes into a different namespace:
python linker.py yeast-interactome.txt query-YMR001C.txt chen2004-species.txt -o cdc5 -m idmap-systematic-to-standard.txt
Thus, Linker generates the following more interpretable mapped image:
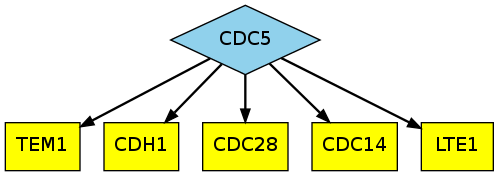
3.) Finally, we are often interested in more than the default k=5 most
probable paths. We can ask for the k=50 most probable paths using the
-k option, which can be provided multiple times.
python linker.py yeast-interactome.txt query-YMR001C.txt chen2004-species.txt -o cdc5 -m idmap-systematic-to-standard.txt -k 10 -k 25 -k 50
When the -k option is given multiple times, a png
image is generated for each unique value of k, and only one
*-pagerank.txt and one *-ksp_*.txt file are created for the
largest value of k. So the previous example generates the following
five files: